Viruses of Protists: Matthias Fischer
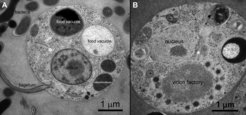
We study very large DNA viruses that infect a group of protozoa called heterotrophic nanoflagellates. Our model organism is Cafeteria roenbergensis, a common marine zooplankton species. The lytic virus that infects this host is named Cafeteria roenbergensis virus (CroV) and belongs to the group of giant viruses. CroV has an icosahedral particle with a diameter of 300 nm that is composed of at least 141 viral proteins and a double-stranded (ds) DNA genome of ≈700,000 bp. We use laboratory-based techniques in combination with comparative genomics and phylogenetic analysis to assign biological functions to unknown proteins, study details of the viral infection cycle, elucidate the genetic relationship to other giant viruses and make predictions about the evolutionary history of this unusual group of viruses.
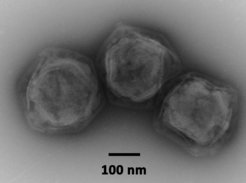

CroV is a cytoplasmic virus, which means that its infection cycle is carried out exclusively in the host cytoplasm with no known nuclear stages. As a prerequisite for this replication mode, CroV encodes its own set of transcription proteins that allow viral gene expression to occur in the cytoplasm. The subcellular compartment, which is created by CroV during infection and where transcription, DNA replication, and particle assembly take place, is called the virion factory. This elaborate viral organelle is highly structured and contains various enzymatic functions. The biochemical complexity of the virion factory is in turn exploited by another group of viruses called virophages.
Virophages are dsDNA viruses with ≈20-30 kb genomes and icosahedral particles with a diameter of ≈75 nm. They infect protists, but cannot replicate themselves unless a specific giant virus co-infects the same host cell. Unlike DNA viruses of comparable size that replicate in the nucleus (e.g. adenoviruses), virophages have evolved to parasitize the cytoplasmic virion factory of giant viruses. Their gene expression depends on the giant virus-encoded transcription machinery. Virophages are true parasites of giant viruses, because the production of progeny giant viruses is inhibited in the presence of a virophage. Host populations infected with a lytic giant virus benefit from virophages and show increased survival rates.
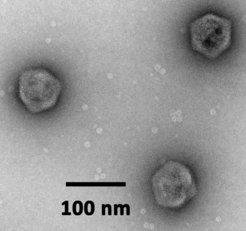
In the Cafeteria-CroV model system, we investigate the infection cycle of the mavirus virophage and its interactions with CroV and the cellular host. In particular, we are interested in how mavirus integrates into the C. roenbergensis genome and how the resulting provirophage is reactivated by CroV infection. We study virus-host dynamics to quantify the effects of mavirus on CroV and the host population, and we explore the diversity of endogenous virophages in protist genomes. Finally, we use computational approaches to study the evolutionary connections between virophages, Maverick/Polinton DNA transposons, and a newly discovered class of endogenous viruses called polintoviruses.


