In the Spotlight: Pump-Probe Photo-Illumination of Biomolecules
Improving structural investigations at X-ray free-electron lasers
Understanding the mechanisms of light-sensitive proteins is highly relevant, as they are key players in life, for example, in vision or photosynthesis. Experiments at X-ray free-electron lasers contribute to the understanding of these proteins through providing high-resolution structural information. Scientists at the Max Planck Institute for Medical Research and French collaborators describe how to conduct those measurements correctly to enable observing protein dynamics closest to their natural reaction. They also point out common pitfalls and mistakes. The brief communication was published in Nature Methods this week.
Sunlight is an essential and ubiquitous element of our environment. Therefore, most organisms have evolved proteins to sense it, make use of its energy or avoid its damaging effects, particularly damage due to UV light. Understanding the molecular mechanisms of these proteins is not only of interest from a basic science point of view, but is also highly relevant to pursuits in medicine (e.g. vision) and technology (artificial photosynthesis, solar cells).
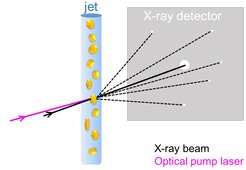
Many relevant light-induced processes occur on the ultrafast timescale, the traditional realm of spectroscopy. Indeed, spectroscopic analysis provides very important insight into the energetic states of the system but it cannot offer high-resolution structural information. With the advent of X-ray free-electron lasers (XFELs), this gap has been bridged: The ultrashort, highly intense X-ray pulse from an XFEL enables high-resolution structural investigations on chemical time-scales of femtoseconds (1 fs = one millionth of a billionth of a second). This allows scientists to follow photon-induced changes in a molecular structure from the very instant of photon absorption onwards. To start the reaction, a very short “pump” pulse of laser light photoexcites the molecule. After a certain time delay the evolving system is probed by an XFEL X-ray pulse, an approach dubbed “pump-probe”. Snapshots collected after different time delays allow one to follow the reaction akin to a flip book (Daumenkino).
Given the unprecedented insight that can be obtained, a number of studies on different light-sensitive systems have now been performed by researchers worldwide. Unfortunately, these studies have all too often been performed using inappropriately high pump laser intensities, exceeding the power of sunlight by factors of billions, resulting in the molecules under investigation being excited multiple times. Such experiments then probe only biologically irrelevant reactions. In their Nature Methods publication, researchers from the Max Planck Institute for Medical Research and from France describe how to plan and conduct pump probe experiments at XFELs correctly such that the desired, naturally occurring biological structural changes can be observed. ”We point out common pitfalls and mistakes, says Marie Grünbein, first author and PhD student in the Department of Biomolecular Mechanisms at the MPI for Medical Research, “this will allow remarkable insight into biological molecules at work, under conditions as similar as possible to nature. If we want to understand these processes, we have to start them correctly in the first place.”
